Integrated analysis of Head and Neck squamous cell carcinoma: A genomic approach
Researcher:Dr. Pawan Kumar Upadhyay
Funding:
DBT-Wellcome Trust India Alliance
Collaborators:
Dr. Sudhir Nair, Tata Memorial Hospital, Mumbai
Tongue cancer is the most predominant form of oral cancer in developed countries with varying incidence in developing countries wherein etiological factors such as chewing betel-quid comprising betel leaf, areca nut and slaked lime is a part of the tradition. While several large-scale genome-sequencing efforts of advanced stage primary oral tumors have been described, systematic efforts to catalogue somatic alterations in tobacco/ nut chewing associated early stage (pT1/2) HPV-negative tongue tumors has been lacking.
Figure: Identification of somatic mutations and DNA copy number changes in HPV-negative early stage TSCC tumors.
In a comprehensive effort to characterize tongue tumors derived from Indian patients:
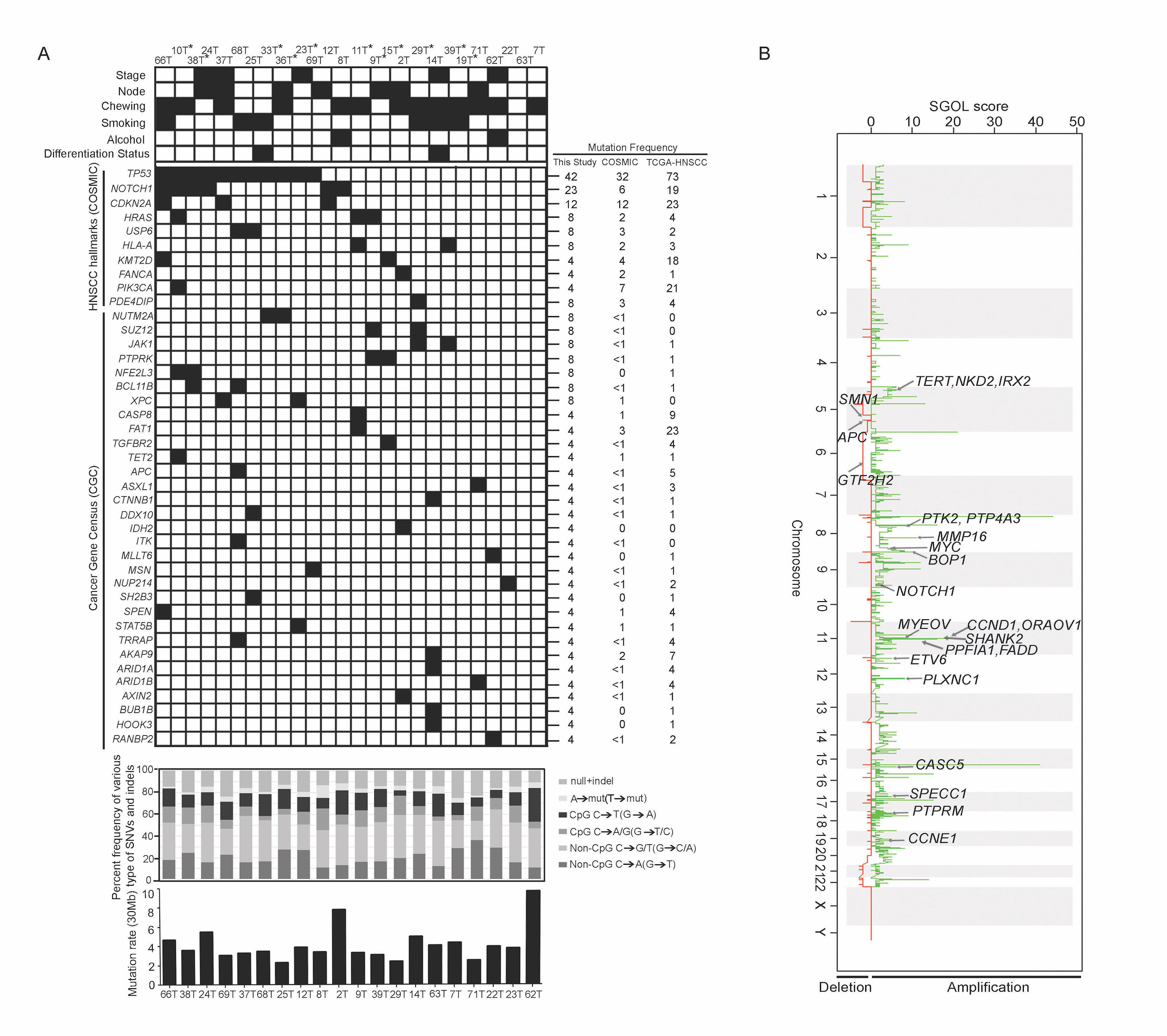
(a) Mutational features of 25 early tongue squamous carcinoma samples (b) Somatic DNA copy number changes identified using Exome sequencing data. Somatic DNA copy number gains and losses were generated using Segments-of-Gain-Or-Loss (SGOL) scores across 23 TSCC patients. Representative amplified and deleted regions are annotated for HNSCC-associated genes and denoted by an arrow. (Oral Oncology. 2017).
Identification and characterization of potential biomarkers to predict metastasis of early stage oral cancer
Researcher:Bhasker Dharavath, Ph.D-SRF
Funding:
DBT-Wellcome Trust India Alliance
Collaborators:
Dr. Sudhir Nair Tata Memorial Hospital, Mumbai
Dr. Anil D’Cruz Tata Memorial Hospital, Mumbai
Dr. Kumar Prabhash, Tata Memorial Hospital, Mumbai
Nodal metastases status among early stage tongue cancer patients (pT1-pT2) plays a decisive role for choice of treatment, wherein about 70% patients may be spared from surgery with accurate prediction of negative pathological lymph node status. Thus, there is an unmet need for identification of prognostic biomarkers to stratify the early stage tongue cancer patients who are likely to develop metastases. To identify genes and miRNAs implicated in early stage metastasis of tongue cancer, we performed whole transcriptome sequencing and microRNA profiling of early stage tongue tumor samples and metastatic lymph nodes. Further, the candidate genes and miRNAs are being validated in extended cohort of oral cancer patient samples, followed by functional characterization of target genes using in vitro and in vivo studies. Preliminary studies suggest that a matrix metalloproteinase gene, MMP10, and a set of 8 miRNAs could play an important role in invasion and metastasis of oral cancer and could act as potential candidate biomarkers to predict nodal metastasis in early tongue cancer patients.
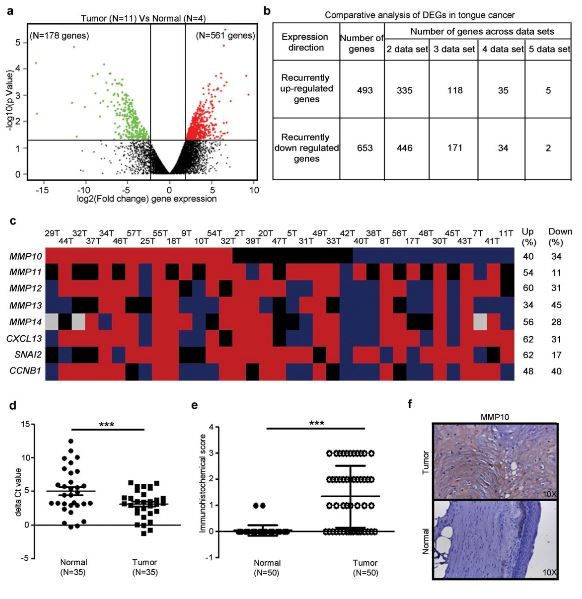
Figure: Differential expression profile of tongue squamous cell carcinoma using mRNA sequencing
and meta-analysis identifies MMP10 up regulation.
Differential expression analysis to identify the distinct gene expression profile of tongue tumors.
(a) Volcano plot representation of differentially expressed in between early tongue tumors and adjacent
normal tongue tissues. The red and green dots denote the up-regulated and down-regulated differentially
expressed genes with P value < 0.05 and fold changes ≥ 2 or ≤ -2 for, respectively.
(b) The tabular representation of a number of genes overlapped in tongue cancer across different studies.
(c) Schematic representation of commonly up-regulated genes qRT-PCR validation in a cohort of 35
paired tongue tumor samples. The Red denotes up-regulation, blue as downregulation, black as basal
expression and gray color; experiment could not be done or results could not be acquired. The ≥2 mean
fold change is for up-regulation, ≤0.5 mean fold change for down-regulation and in between 1.99-0.501
mean fold change as a basal level expression compared to the adjacent normal tissue sample.
(d) qRT-PCR analysis of MMP10 transcript expression in early tongue tumors. Dot plot representation of
MMP10 transcript ΔCt value distribution and its significance between normal and tumors tongue tissue
sample (n=35). Each dot represents the average normalized ΔCt value of MMP10 in a single sample.
Median with interquartile range is shown for MMP10 for normal and tumor samples.
(e) Dot plot representation of immunohistochemical score of MMP10 expression in tongue tumors and
adjacent normal tissues (n=50). Each dot represents that final IHC score for each sample and median with
interquartile range is shown. Median with interquartile range is shown for MMP10 protein expression in
normal and tumor samples.
(f) Representative IHC stained photomicrographs tongue tumors and paired normal samples are shown.
The brown color indicates positive expression of MMP10 protein. The P-value was calculated by Mann-
Whitney U test using GraphPad Prism 5 program and p value ≤0.05 was considered as a threshold for
statistical significance. P value is denoted as ***; P < 0.0001[5].
Publications: