
PCR based concatenation of KRAS and EGFR exons
Clinical collaborator: Dr. Kumar Prabhash (TMH)EGFR and KRAS mutations occur at variant frequencies ranging from 10-60% across different ethnic populations. Genotype-based stratification of patients for presence of KRAS and EGFR mutation has immense clinical implication in potentially precluding EGFR TKI therapy from some patients as is in current practice in lung and colorectal cancer. Of several different testing methods available, PCR followed by directed sequencing and amplification refractory mutation system (ARMS) are the two most commonly used diagnostic methods. While ARMS methodology has several advantages of higher sensitivity but is cost prohibitive, especially for regional cancer care centers in the third world countries. Compared to ARMS, directed sequencing is relatively inexpensive, widely used but more cumbersome to perform. Moreover, with limiting amount of genomic DNA from clinical FFPE specimens or fine biopsies of lung tumors, either of the methodology gets challenging to perform. Furthermore, neither of these two methodologies in its current form can be utilized at high throughput mode to determine complete spectrum of EGFR and KRAS mutations using targeted next generation sequencing.
In this study, we report a novel and cost effective single multiplex-PCR based method, CRE, for co-amplification of five KRAS and EGFR exons followed by concatenation of the PCR product as a single fragment—an improvised protocol for direct sequencing. CRE is a robust methodology that has a potential to be routinely used in clinical diagnostics with reduced variability, cost and turnaround time requiring minimal amount of template DNA extracted from FFPE or fresh frozen tumor samples. As the limitation of the CRE strategy is defined by the sensitivity and resolution of the sequencing methodology adopted, concatenated PCR products from multiple individuals—each tagged with unique bar code sequence—can be pooled and deep sequenced using a NGS platform. In nutshell, this is a multiplex PCR based concatenation methodology report with application to cancer diagnostics, along with potential for high throught based profiling.
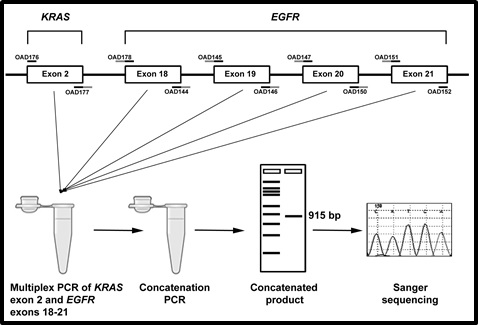
[Schematic representation of CRE: Concatenation of KRAS and EGFR exons: Workflow for CRE methodology. KRAS and EGFR primers are shown along with complementary tail overhangs that primes with consecutive exons in an ordered manner. PCR products amplified with a cocktail of primers for KRAS and EGFR exons in a single multiplex reaction is transferred to a fresh tube and concatenated in a separate reaction.]